|
1 Comment
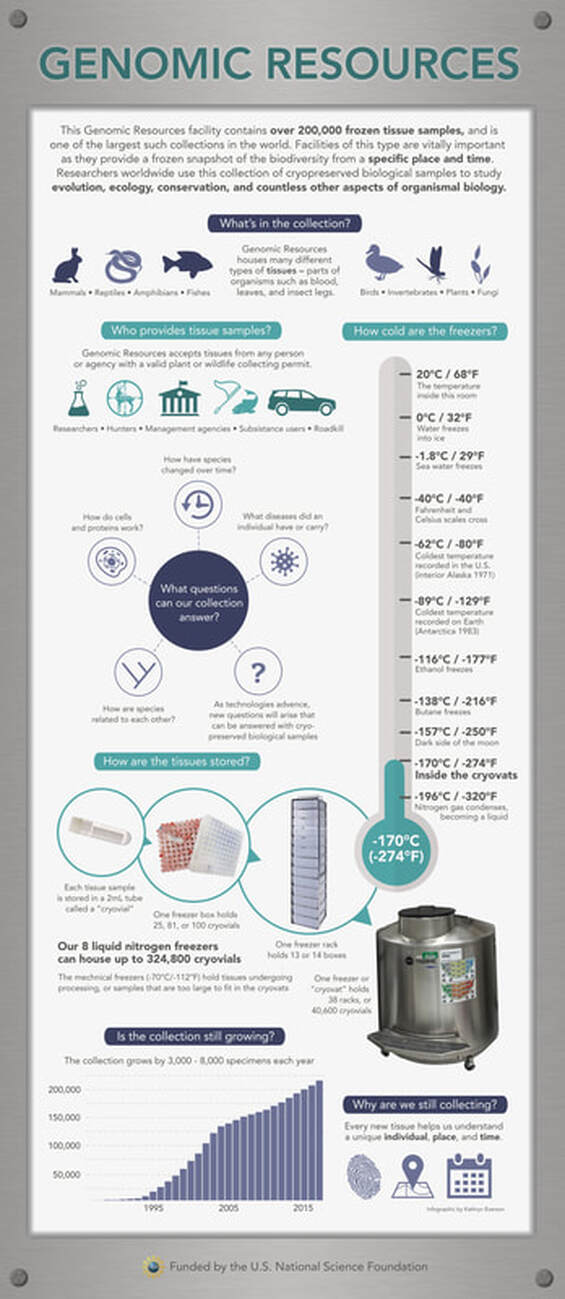
From the archives: A poster for the genomic resources collection at the University of Alaska Museum1/14/2021 Back when I was working in the University of Alaska Museum (or as it's known to the public, the "Museum of the North") I was asked to create a poster to hang on a tall, skinny section of wall outside the genomic resources collection. The goal was to provide information for visitors of the public to understand what the genomic resources collection actually is! See the poster below (and at the Museum of the North, if you ever take a tour!)
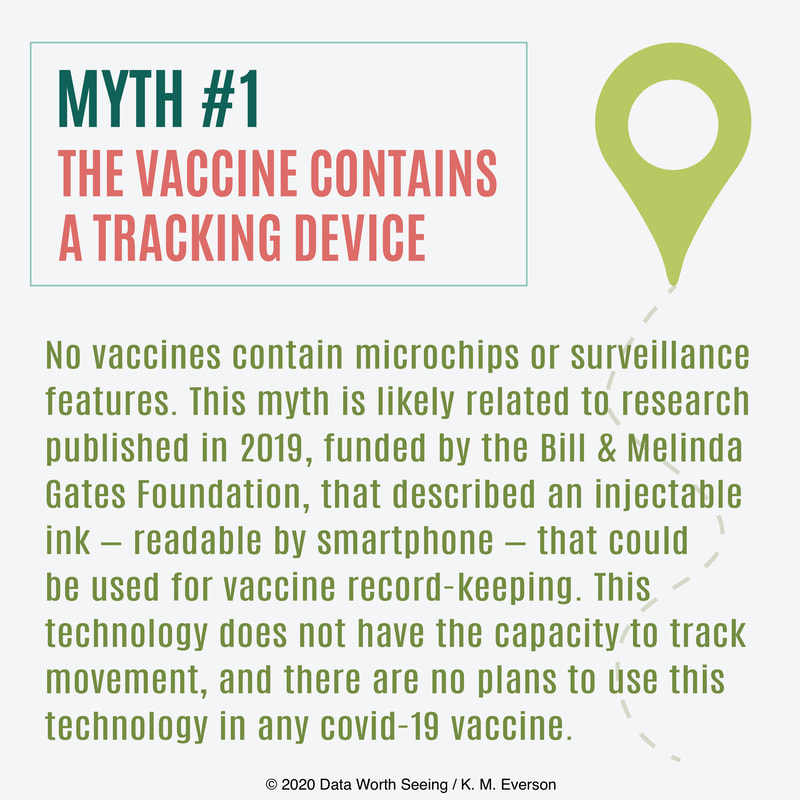
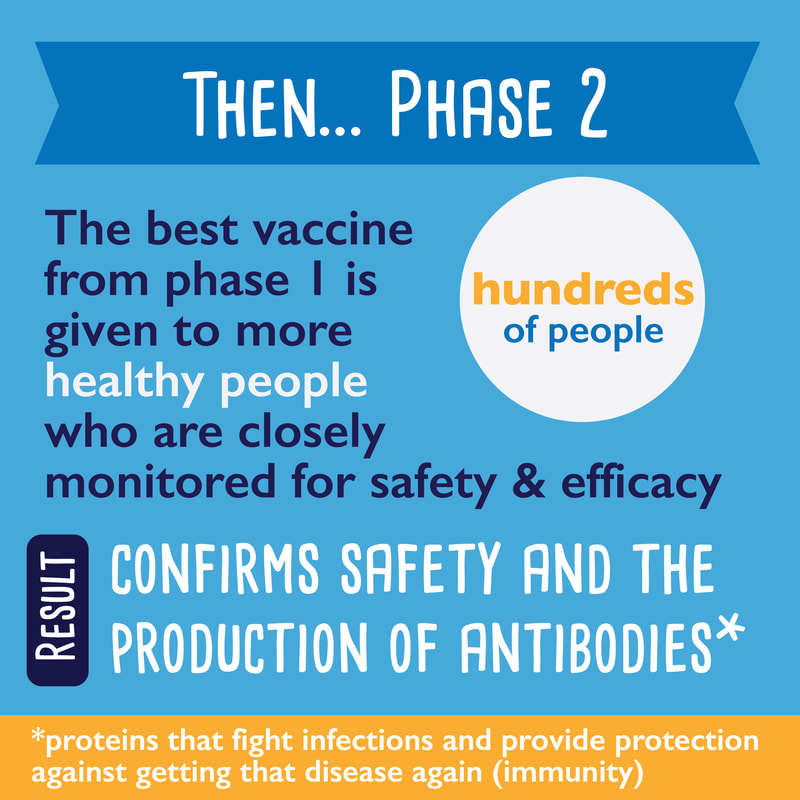
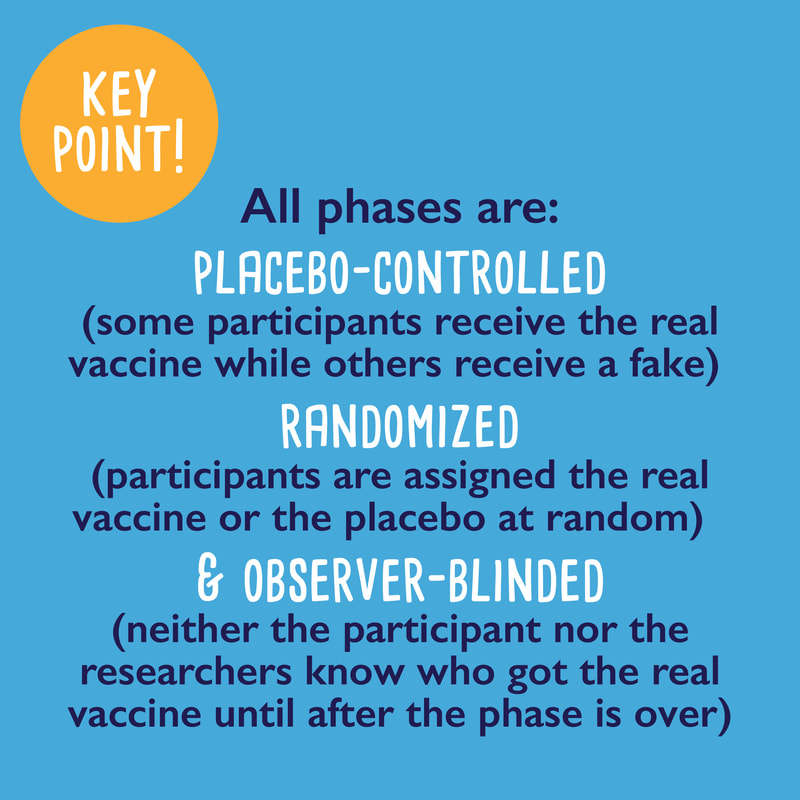
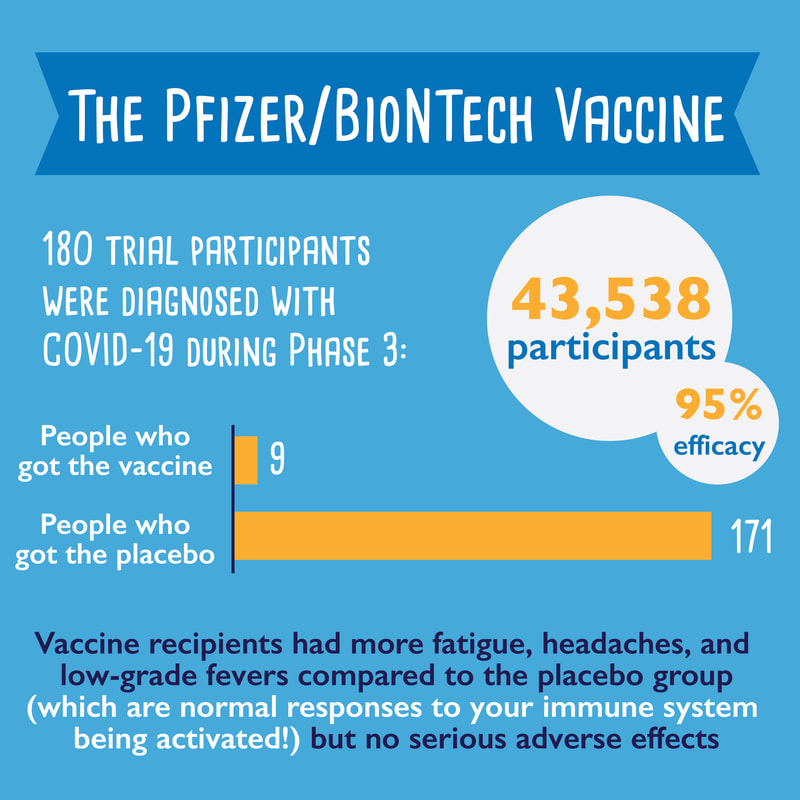
Part 1 of a 3-part series about coronavirus vaccines: What are the different vaccine types?1/14/2021 I've noticed a lot of misinformation going around town about coronavirus vaccines... are they safe? Do they work? How are they different from other vaccines? Here is part 1 of a 3-part series where I try to answer some of those questions:
|
AuthorI'm Katie Everson, a postdoc at the University of Kentucky. Archives
January 2021
Categories
All
|
| contact me: |
























 RSS Feed
RSS Feed
